From the Journals: MCP
A deep learning approach to phosphoproteomics. Untangling complex proteomics mass spec data. Read about papers on these topics recently published in the journal Molecular & Cellular Proteomics.
A deep learning approach to phosphoproteomics
The post-translational modification known as phosphorylation regulates numerous cellular functions, including cell growth, movement and metabolism. Phosphoproteomics, the study of phosphorylated proteins, offers valuable insights into dynamic changes in signaling pathways mediated by phosphorylation. Phosphoproteomics relies on mass spectrometry and computational methods to detect phosphorylated sites within a protein sequence. However, accurate detection is challenging due to the transient nature and low abundance of phosphorylated peptides relative to the proteome. Moreover, different computational methods yield varying outcomes, thus undermining confidence in the results.
In a recent study published in Molecular & Cellular Proteomics, Xinpei Yi and colleagues from Baylor College of Medicine and the Liver Cancer Institute at Fudan University employed deep learning to enhance phosphoproteomics accuracy. Deep learning is an artificial intelligence method that discerns patterns from vast amounts of unstructured data using neural networks.
Yi and collaborators used a multifaceted approach that included deep-learning algorithms for predicting peptide retention time and fragment ion intensity, a statistical scoring algorithm that integrates these deep-learning predictions for determining the probability of a site being phosphorylated, and a machine learning algorithm that also integrates the deep learning predictions for identifying peptides based on theoretical spectra. Compared to standalone algorithms, this integrative approach, named DeepRescore2, identified more phosphorylated peptides in synthetic and biological data sets from normal and liver cancer tissues.
Accurately identifying phosphopeptides can aid in precise inference of the activity of kinases, which drive phosphorylation. The authors used DeepRescore2 to infer activity of epidermal growth factor receptor kinase, a biomarker of liver cancer. DeepRescore2 correctly inferred high kinase activity in liver cancer samples from patients with poor prognoses. The combinatorial deep learning approach used in the study enables accurate profiling of phosphoproteins and may serve as a prognostic tool. Combining DeepRescore2 with other algorithms enables more precise detection of phosphorylation activity, thus paving the way for biomarker-guided cancer therapies.
Untangling complex proteomics mass spec data
Data-independent acquisition, or DIA, is a popular technique used to analyze proteomes by mass spectrometry, or MS. In DIA, all detectable ions from a proteomics sample are iteratively analyzed, much like observing a crowded street one angle at a time. Despite the broad coverage and sensitivity DIA provides, its application is limited due to the difficulty in discerning similar molecules with overlapping signals in a complex sample.
In a recent study published in the journal Molecular & Cellular Proteomics, Sophia Steigerwald at the Max Planck Institute of Biochemistry and colleagues combined DIA with an advanced signal processing method, the phase–constrained spectrum deconvolution method, or ΦSDM, to tackle the challenge of complex spectra. The ΦSDM method imposes mathematical constraints based on the phase information of peptide ions to untangle overlapping signals within a mass spectrum. The authors tested the ΦSDM–DIA approach using HeLa cells.
By implementing ΦSDM signal processing on additional graphics processing units, the researchers quickly achieved a higher signal-to-noise ratio and 15% more peptide coverage compared to conventional methods, particularly in samples with short gradient times. Thus, advanced analytical techniques such as ΦSDM–DIA may help unravel unknown cellular pathways and events driven by complex proteomes or transient proteins without compromising on speed, quality and accuracy.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Flipping lipids and slime molds
A dull first job nearly pushed JBC associate editor Todd Graham out of science. Then a slime mold project changed his path. Now, he studies membrane biology and reflects on discovery, persistence and mentoring through uncertainty.
How smelling death alters worm behavior
Researchers have found that the roundworm C. elegans can smell death, and it changes how the worms behave, reproduce and age.

A chance encounter with the lab
Payton Stevens never planned to become a pancreatic cancer researcher. A temporary job set him on a path from rural Kentucky to leading research on Wnt signaling and metastasis, where he now pairs discovery with mentorship and science advocacy.

Light-activated small molecule could transform eye infection treatment
Contact lenses raise the risk of infectious keratitis, a leading cause of blindness worldwide. A biotech company is commercializing a light-activated therapy using a ROS-generating molecule to rapidly kill microbes in the cornea to preserve vision.

The molecular orchestra of memory
Calcium, calmodulin and calcium/calmodulin-dependent kinase II form a molecular axis that turns fleeting neural activity into lasting memories. New research shows how memories are stabilized, and possibly even protected or repaired.
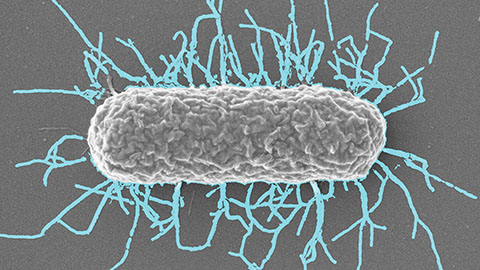
Differences in pili structure modulate bacterial behavior
Researchers demonstrate how small changes in the structure of hair-like protein appendages can affect the behavior of Acinetobacter bacteria.

