Illuminating the dark serum immunoglobulome
Most of us know that the symptoms of COVID-19 can range from nothing to catastrophic systemic failure, depending on the individual. A recent study published in the journal Molecular & Cellular Proteomics showed that patient antibody responses to the SARS-CoV-2 virus that causes COVID-19 are highly personalized and affected by the disease course and treatment.
Human blood is packed full of cells, proteins, sugars and dissolved gases. However, the most common proteins in serum are immunoglobulins, or antibodies. Immunoglobulins are glycoproteins produced by adaptive immune cells, and they act by recognizing and binding to antigens derived from pathogens, such as bacteria or viruses, thereby, initiating the pathogens’ destruction.
Despite their abundance and importance to human health, antibodies are often under-studied because their immense sequence diversity presents a technical challenge to researchers.
“Our group is a mass spectrometry group and not an immunology group,” Albert Bondt, project manager at University of Utrecht, Netherlands, and lead author of the study, said. “We come from a background of studying individual proteins and their finest details. From that background, we don't look immediately at the whole population. But we look at every individual, and, even within the individual, we look at individual molecules.”
The Utrecht researchers developed a method to study what they call the “dark” serum proteome. This mysterious group comprises proteins that scientists think should exist but haven’t found, proteins that can be constructed and modified in different ways and proteins with unknown structures and functions.
“There are potentially billions of different antibodies in the blood,” Bondt said. “We found that you can actually only detect a few, maybe 100, that are quite abundant. But there are likely still billions that aren’t detectable at the moment.”
To begin illuminating those billions of unknown antibodies, the group used mass spectrometry–based methods to sequence individual antibodies of two subtypes, IgG1 and IgA, over time in a cohort of 17 COVID-19 patients.
“We capture the antibodies, and then we remove the part that introduces a lot of variety (Fc region), so we can look at the part that binds the antigen and really matters (Fab region),” Bondt said. “Then, we get a picture for every individual of all the antibodies that are present in the blood.”
These antibody clonal profiles were used to examine how each patient reacted to a severe SARS-CoV-2 infection.
“For the sample prep, we assumed the average values that every textbook says someone should have for these types of immunoglobulin,” Bondt said. “But when we look at our data, we actually see that there's a huge variation between individuals. So, if the textbook says you should have five milligrams of IgG1 per milliliter of blood, that can be true. But for some patients, their IgG1 levels are 0.1 milligrams (per milliliter), and for others, they have five times as much.”
Bondt said he and his team were surprised to find that each patient had unique ratios of IgG1 and IgA that changed over time.
“Textbooks would state that IgG1 is always more abundant than IgA (in the blood),” Bondt said. “Well, according to our data, that is not the case per se. There are many individuals with more IgA than IgG1. These kinds of things may seem obvious, but they are often ignored.”
Bondt said his team’s mass spectrometry–based approach can probe much deeper into antibody repertoires than previously used methods. Therefore, the approach could be used to monitor patient therapeutic responses over time, which would allow physicians to modify or discontinue treatment when a desired antibody level is achieved.
“If there were to be another pandemic, for example, I think we are way more prepared thanks to this technology,” Bondt said. “We would be able to identify and provide therapeutic antibodies at a much faster pace.”
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Light-activated small molecule could transform eye infection treatment
Contact lenses raise the risk of infectious keratitis, a leading cause of blindness worldwide. A biotech company is commercializing a light-activated therapy using a ROS-generating molecule to rapidly kill microbes in the cornea to preserve vision.

The molecular orchestra of memory
Calcium, calmodulin and calcium/calmodulin-dependent kinase II form a molecular axis that turns fleeting neural activity into lasting memories. New research shows how memories are stabilized, and possibly even protected or repaired.
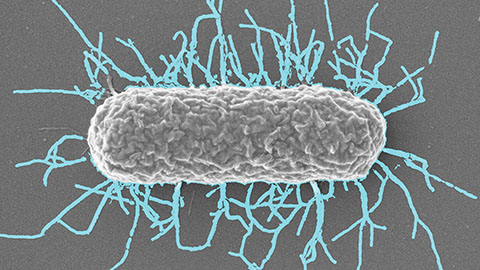
Differences in pili structure modulate bacterial behavior
Researchers demonstrate how small changes in the structure of hair-like protein appendages can affect the behavior of Acinetobacter bacteria.

Cholesterol regulatory genes predict liver transplant outcomes
Researchers identify a link between cholesterol-regulating genes and liver transplant success, which could improve donor screening and patient outcomes.

Lipid signatures for a rare neurological disorder
Researchers find distinct lipid patterns linked to a rare autoimmune neurological disorder, offering hope for effective targeted therapies for patients.
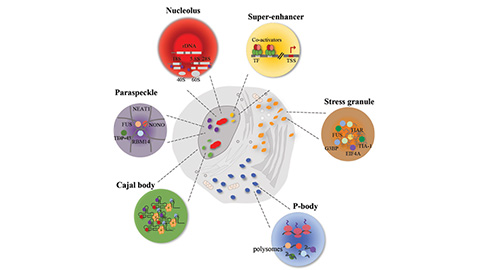
Disease-linked mutations disrupt protein phase behavior
Researchers find that pathogenic missense mutations are enriched threefold in phrase-separating intrinsically disordered regions of proteins.

