MCP: Proteogenomics researchers find causes of immune disease
Routine clinical sequencing has given doctors unprecedented insight into genetic disorders. However, genomics fails to diagnose up to half of patients who are tested. That’s the problem that scientists at universities in Munich and Berlin tackled in a recent study in the journal Molecular & Cellular Proteomics. With samples from patients in four countries and a novel database on the neutrophil proteome, Christoph Klein and colleagues diagnosed two children with severe congenital neutropenia using mass spectrometry-based proteomics when typical sequencing had failed.
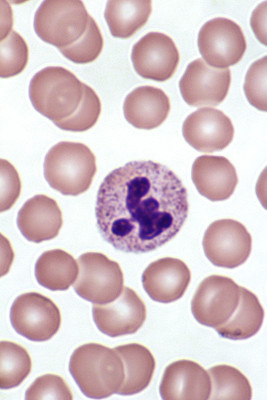 Neutrophils, like the one in the center of this photo, are loaded with granules full of proteases that make them difficult to study. Guy Waterval /Wikimedia Commons
Neutrophils, like the one in the center of this photo, are loaded with granules full of proteases that make them difficult to study. Guy Waterval /Wikimedia Commons
“There are very few examples of people who use multiple omics to investigate rare diseases … (but) I think this is the future of personalized medicine,” said Klein, a physician and director of the Children’s Hospital of the University of Munich.
The patients’ disease affects their neutrophils, white blood cells packed with toxic proteins to deploy against bacteria. When neutrophil development is disrupted, which Klein estimates happens to 1 in 200,000 newborns, every bacterial or fungal infection can become a life-threatening medical emergency.
Klein’s lab has studied rare genetic causes of neutropenia for years, but proteomics was a new field for the group. Postdoctoral researcher Sebastian Hesse met proteogenomics expert Juri Rappsilber at a conference, sparking a collaboration to study the proteome and transcriptome of neutrophil granulocytes.
Neutrophils are post-mitotic and very fragile, which makes studying them a challenge.
“You can think of them as suicide bombers,” Klein said, explaining that the cells are full of granules loaded with proteases that make retrieving other proteins a challenge. Hesse painstakingly developed a protocol to collect intact proteins and mRNA from healthy neutrophils.
Using mass spectrometry, scientists led by co-first author Piotr Grabowski in the Rappsilber lab at the Technical University of Berlin analyzed the cells’ proteome. When they added transcriptomic data, they found strikingly little correlation with the proteome, so they chose to focus on protein in patient samples.
Next, Hesse collected neutrophils from 16 patients with congenital neutropenia. Some were in Germany; to find others, he had to travel.
“These patients are from various parts of the world — that’s another unique challenge of working with rare diseases,” Klein said. “Sebastian flew to Iran and Turkey, in a collaborative effort with the pediatric hospitals there.”
Back at home, after processing and freezing the samples, Hesse handed them off to Grabowski; the proteomics scientists repeated the analyses to see what proteins had changed in the patients’ blood.
The team used abnormal protein profiles to guide the diagnosis of two patients with inconclusive exome sequencing results.
In one child’s case, a pseudogene made it difficult to identify mutations in the protein-coding gene; in the second, incomplete coverage by exome sequencing had missed a key point mutation. Data on protein abundance in each patient led the researchers to run more specific genetic tests that proved conclusive.
“This highlights (that) even if you have highly controlled pipelines for genetic studies, there’s always a risk that you are not 100 percent correct,” Klein said.
The researchers did not set out to make diagnoses from patient proteomes, but the study highlights the value proteomics data can add.
“Cellular proteome studies are not in routine clinical use at this point,” Klein said. “But … I think there will be huge potential for proteome analysis at a very low cost down the road.”
The team plans to expand its studies to other patients with immune deficiencies, looking for new genetic mechanisms of disease.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles
Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.
Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.
Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.
Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.
Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

