Quantifying how proteins in microbe and host interact
Antimicrobial resistance is a growing problem around the world. When microbes such as bacteria, viruses, parasites and fungi stop responding to antimicrobial medicines, most researchers respond by working to develop new vaccines. However, when host proteins bind to bacterial proteins, that binding masks some dominant antigenic sites on the pathogen. Because of this, traditional vaccines have seen poor antibody responses.
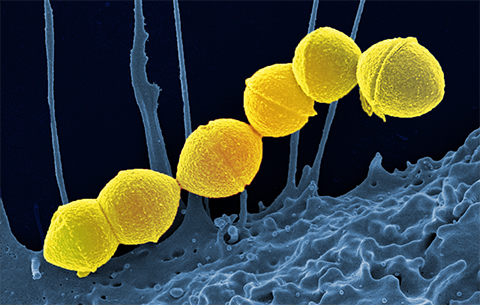
In a recent study published in the journal Molecular & Cellular Proteomics, researchers at Lund University used affinity purification combined with quantitative mass spectrometry, or AP-MS, techniques to investigate if these bacteria and host protein–protein interaction networks could compromise the development of protective antibody responses; they used Streptococcus pyogenes, a bacteria that causes infections in humans,as a model system.
Johan Malmström was the corresponding author of this study. “To develop better vaccines, we need new methods and a better understanding of the antibody responses that develop in immune individuals,” he said.
Streptococcus pyogenesbelongs to Group A Streptococcus, or GAS, which affects about 18 million people annually, and claims over 500,000 lives, so an urgent need exists for a vaccine against GAS.
In previous work, Malmström’s lab has shown that GAS forms extensive protein interaction networks with human proteins, so they wanted to analyze the association between the antigen-specific antibody responses and the host protein–bacterial protein interactions. They also wanted to understand if antigen-specific antibody responses could affect these protein networks to understand antibody repertoire (the entire set of antibodies produced in an individual), ultimately aiding the development of effective vaccines against GAS.
In this study, the researchers immunized mice with the conserved M1 protein from GAS and showed that mouse plasma protein forms similar protein networks with the bacterial M1 protein as those seen in humans. Using AP-MS, they quantified the competition among antibodies and protein interaction networks.
While the team found that the antigen targeted by the anti-M1 antibodies affected sites outside the protein interaction surfaces and could not outcompete these interactions, they also discovered in this study that regions in bacterial proteins within protein-interaction interfaces are more challenging to develop antibodies against, highlighting the need to quantify and assess these interactions.
Several bacterial pathogens use their surface proteins to interact with the host proteins to dodge the immune system and mask their dominant epitopes.
The experimental strategy developed in this study can measure the level of competition between antibodies and host plasma proteins at the same bacterial sites, thereby dictating the course of host immune responses. This method could be broadly applicable to any type of bacterial protein.
“I think that the most efficient way forward would be to build networks of scientists with different research skills and meet around the same problem,” Malmström said.
Read more about this group’s research here.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles
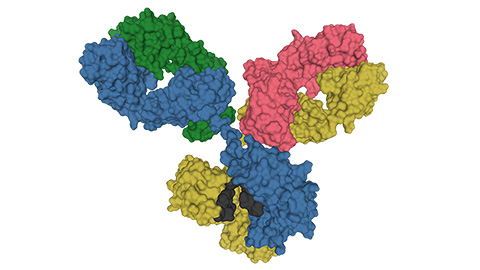
Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.
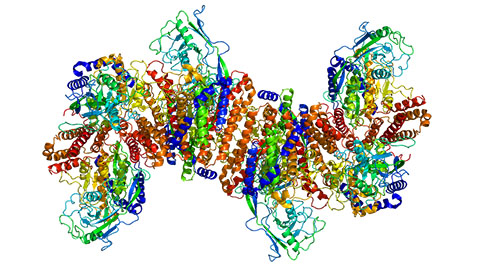
Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.
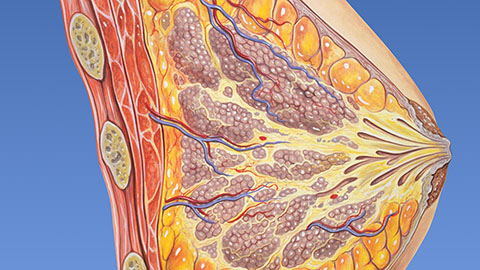
Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

Flipping lipids and slime molds
A dull first job nearly pushed JBC associate editor Todd Graham out of science. Then a slime mold project changed his path. Now, he studies membrane biology and reflects on discovery, persistence and mentoring through uncertainty.
How smelling death alters worm behavior
Researchers have found that the roundworm C. elegans can smell death, and it changes how the worms behave, reproduce and age.

