
CALeDNA: Tracking biodiversity at the molecular level
An estimated 9 million kinds of plants, animals, protists and fungi live on Earth. Biodiversity loss is a major problem associated with human-made climate change. The current global rate of extinction is predicted to be nearly 100 times the baseline rate. But measuring those extinctions, or even accurately measuring the organisms that make up an ecosystem, is a challenge, since biodiversity assessment must include species from communities that cannot be observed by eye.

as frozen collections.
CALeDNA (pronounced cal-ee-DNA) is a partnership of scientists, universities and nonprofits focused on measuring ecosystems’ biodiversity and how ecosystems change over space and time. To do this, they isolate environmental DNA, or eDNA — which organisms shed into the ecosystem — from water, soil or sediment samples. They use a method called metabarcoding to amplify and sequence specific genetic loci including ribosomes, internal transcribed spacers, and cytochromes and then identify a list of species with eDNA in each sample. Sample collections are archived at University of California labs as a resource for future projects.
The program emphasizes community involvement. CALeDNA organizes bioblitz events where community members and local organizations come together in the field to collect samples, photographs and geolocation data. The data from these projects are made publicly available through the CALeDNA website and data exploration web tool.
Rachel Meyer is the CALeDNA director at the University of California, Santa Cruz. She is also an adjunct assistant professor and part of the UCSC Paleogenomics Lab led by Beth Shapiro and Ed Green. I sat down with Rachel to learn what CALeDNA was all about.
What ideas led to this program? What challenges are you facing?
The idea was to get a biodiversity signature from DNA in the environment that could be systematically collected around the globe to balance out biodiversity observations and improve predictive models. The proof of concept came out of work with ancient DNA. Eske Willerslev and collaborators including Beth Shapiro were the first to show that you could extract ancient and modern DNA from environmental samples, be it ice cores, soil and sediment, or bones. Getting biodiversity data from eDNA was working, and people were excited about it in Europe but not as much in the States.
One of the reasons people have been reluctant to adopt eDNA in monitoring and modeling is people are skeptical about false positives and negatives. Another reason is we just don’t know how to put eDNA together with other data types to strengthen our analysis. When we started trying to analyze eDNA, the tools out there didn’t meet our needs. We had to string together a lot of different software. Then, when we finally got results of species in different samples, we saw different species from what people recorded with traditional observations like photos. When you get an answer you don’t expect, is that false? Or is there an eDNA realm we can’t see? We have been answering these questions slowly with pilot studies.
Fascinating! What led you to your position as the CALeDNA program director?
I went to a Department of Defense conference on environmental conflict in 2016, and they were talking about predicting environmental conditions that cause conflict around the globe. A lot of this is based around water. None of their models included biodiversity; they said it was too hard. I felt like I could contribute to this work, and this led me to start looking for biodiversity research jobs.
What are your goals, and how close do you think you are to achieving them?

We want to be able to measure biodiversity accurately and measure how it changes, and we want to integrate eDNA data with as many other kinds of data as we can.
I think we are still 10 years away from making this standard, but we are getting out of perpetual pilot mode. We are repeating our pipeline now — not building it anymore — so we can start to stack up the pilot projects and glean patterns. And we have recently received financial support to start co-analyzing eDNA with remote sensing climate data from NASA, or functional chemical diversity data, hydrological or biogeochemical data, and anything else under the sun, so we can make sense of changes in biodiversity patterns. We need to broaden our collaboration community, and that takes time.
What do you mean by remote sensing and chemical diversity data?
Satellites can detect related species because they produce related spectral signatures, so you can map the community population of, for example, a forest canopy from space. You can detect stress responses from space. With models, you can map underground fungi from space. This is where we need to engage more with biochemists and molecular biologists. Let’s ask questions about the rules of life, evolution and ecology and the missing metabolic or cellular links!
Wow! So what is the rate-limiting step to integrating all this data?
Honestly, it’s collecting reference sequence data for species and understanding the different things we can measure from our environmental collections. 70% of DNA sequences from our data sets will be tossed because we don’t know what species they are.
There are more interesting questions we want to get into. We track taxa, but what about genes? What about the proteins and their conformation or responses to the environment? We are building better analysis tools and infrastructure to ask these kinds of questions, and we hope researchers will play with the nearly 5,000 samples we have already archived.

Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles
Mitochondria shape kidney cell function
Researchers at the University of Washington, Seattle present the first quantitative comparison of mitochondrial interactomes between two epithelial cell types in the kidney.
Long-chain polyunsaturated fatty acids linked to postoperative delirium risk
Researchers show that altered lipid metabolism may contribute to postoperative delirium, a condition linked to increased risk for long-term cognitive decline. The study explores potential disease mechanisms, which have yet to be understood.
Glycosylation patterns across antibody isotypes distinguish tuberculosis states
Researchers at Taipei Medical University present the first site-specific glycosylation analysis of immunoglobulins in elderly tuberculosis patients.

Blood glycome possibly predicts lifespan
Researchers at the University of Santiago de Compostela show that total serum N-glycome can predict mortality independent of traditional risk factors.
Building a better model for drug delivery across the blood–brain barrier
Industry and academic scientists collaborated to develop a rat with humanized iron-transport receptors, enabling research into iron homeostasis and drugs that cross the brain’s barrier.
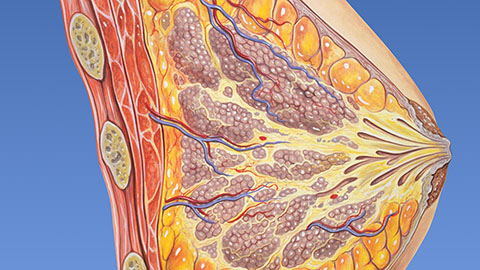
Fat synthesis enzyme crucial for milk fat and newborn growth
Researchers found that a deficiency of the fatty acid synthesis enzyme stearoyl-CoA desaturase-1 reduced mammary gland function during lactation and caused low birth weight in newborns that were fed milk from enzyme-deficient glands.

