The proteome of the cave bear
As a rule, it takes a genome to interpret a proteome.
A genome database gives the range of possible proteins that a sample is expected to contain, allowing a computer program to match short peptide fragments from the raw data to the full-length proteins they came from. The genome is like a picture showing how a jigsaw puzzle will look when it’s finished — and each peptide is a single tiny piece of the puzzle.

Richard Johnson, a staff scientist at the University of Washington’s department of genome sciences, has spent nearly three decades working with no picture. Before genomes were assembled and available, he became an expert in de novo peptide sequencing, piecing together the overlapping puzzle pieces from mass spectra to determine the amino acid sequence of proteins.
That ability has been coming in handy recently since Johnson started seeing more requests for environmental proteomics and other exotic analyses.
“I sit next to an oceanographer, and she does these proteomics analyses on strange samples, like glacial meltwater and seawater,” he said. “Those are cases where it’s really difficult to decide what database to even search.”
To annotate a sample from a human, a researcher can use a human genome database. But a tablespoon of ocean water or glacial runoff is likely to contain a complex community of microbes. So which genome databases should the researcher survey? Usually, researchers solve this problem by sequencing as much DNA as they can from a sample and using the result, a metagenome, to guide protein identification.
But even with a metagenome, sometimes the proteins observed in a proteomics experiment just don’t match the given reference database. “I came up with a metric that can tell you whether the protein sequence database is any good for interpreting your mass spectrometry data,” Johnson said.
The technique, which Johnson and colleagues recently published in Molecular & Cellular Proteomics, can be used to solve related problems, such as proteomic analysis of an animal whose genome has not been sequenced. “You typically use a sequence database from a closely related species and hope that the sequences did not diverge too much,” Johnson said. “Sometimes that hope is warranted, and other times it’s not.”
Johnson has used this approach to study the makeup of electrosensory organs in electric fish.
A third potential application is for analysis of very old but not fossilized tissues — those that come from extinct species, such as a vial of powdered cave bear bone that Johnson’s team obtained. Extinct species very rarely have a genome assembled, and the close-cousin conundrum is compounded by slow biochemical changes to proteins that happen over thousands of years.
But the approach doesn’t solve every problem. Johnson said, “Using this quality metric tells you how good or bad a sequence database is. But it won’t tell you what to do about it if it’s bad.”
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Flipping lipids and slime molds
A dull first job nearly pushed JBC associate editor Todd Graham out of science. Then a slime mold project changed his path. Now, he studies membrane biology and reflects on discovery, persistence and mentoring through uncertainty.
How smelling death alters worm behavior
Researchers have found that the roundworm C. elegans can smell death, and it changes how the worms behave, reproduce and age.

A chance encounter with the lab
Payton Stevens never planned to become a pancreatic cancer researcher. A temporary job set him on a path from rural Kentucky to leading research on Wnt signaling and metastasis, where he now pairs discovery with mentorship and science advocacy.

Light-activated small molecule could transform eye infection treatment
Contact lenses raise the risk of infectious keratitis, a leading cause of blindness worldwide. A biotech company is commercializing a light-activated therapy using a ROS-generating molecule to rapidly kill microbes in the cornea to preserve vision.

The molecular orchestra of memory
Calcium, calmodulin and calcium/calmodulin-dependent kinase II form a molecular axis that turns fleeting neural activity into lasting memories. New research shows how memories are stabilized, and possibly even protected or repaired.
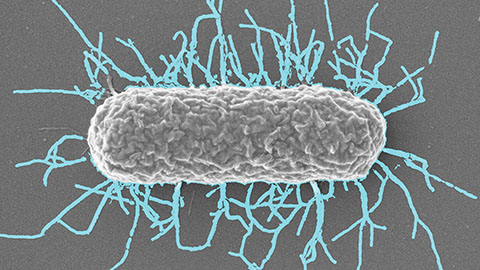
Differences in pili structure modulate bacterial behavior
Researchers demonstrate how small changes in the structure of hair-like protein appendages can affect the behavior of Acinetobacter bacteria.

