A novel technique maps antibodies’ epitope specificities
According to one count, some 5 million commercial antibodies exist, used to target, detect or destroy in contexts from research labs to patients' bodies. Raised in goat, rat, mouse, rabbit, llama and even chicken against fragments of a menagerie of proteomes, these antibodies bind with high affinity to their mostly protein targets.

Determining the portion of a protein surface to which an antibody binds is called epitope mapping. The gold standard approach is to determine the structure of an antibody-target complex. However, structural work can be difficult and may take months per antibody. Researchers also use peptide display technologies, presenting a library of protein fragments on a microarray or the surface of E. coli and finding out which ones the antibody binds to through microarray scanning or flow cytometry.
In a recent article in the journal Molecular & Cellular Proteomics, researchers in the labs of Xiaodong Zhao, Hua Li and Sheng-ce Tao at Shanghai Jiao Tong University introduced a new, faster method for epitope mapping. Using the new technique, they write, "One technician is able to map the linear epitopes of more than 200 antibodies in one month at an affordable cost."
The approach starts with a random phage display library, a collection of 109 peptides with random sequences, each 12 amino acids in length, expressed on the surfaces of phages. (Although other researchers previously have used phage display for epitope mapping, they mostly have presented variants on a known target protein rather than random peptides.)
"The beauty of phage display is the protein sequence and the coding sequence are linked together," Tao said. Therefore, after biochemically extracting phages that bind to an antibody of interest, the team quickly could identify these phages through next-generation sequencing. By comparing all of the phage peptides that bound to each antibody, they detected common protein motifs. This let them establish a consensus binding sequence for each antibody — without any preconceptions about what it would be.
Sometimes the epitope they determined matched the protein or peptide against which the antibody was raised. In other cases, the consensus epitope sequence was not found in the protein an antibody was raised against. According to Tao, this makes sense when you think about antibodies and their targets binding in three dimensions.
"How antibodies are generated is not always based on the linear sequence," Tao said. "Sometimes it's based on conformational structure."
Because of the nooks and crannies folded into any protein's structure, amino acids that are not adjacent in its linear structure often are found next to each other on its surface. While the technique is quite useful for finding linear epitopes, the authors said, it cannot detect conformational epitopes; they hope to find new computational approaches that will bridge the gap.
Still, the research has clear scientific and commercial potential. Since the paper was published, Tao said, the team has improved the speed and efficiency of their technique. The lab has struck up a partnership with a reagent supply company to map the epitopes of 10,000 of its antibodies and anticipates using the resulting data set to learn more about the relationships among antibody, antigen and epitope. They're also considering other applications, such as identifying potential off-target binders that could cause unexpected side effects in antibody-based drugs.
Enjoy reading ASBMB Today?
Become a member to receive the print edition four times a year and the digital edition monthly.
Learn moreGet the latest from ASBMB Today
Enter your email address, and we’ll send you a weekly email with recent articles, interviews and more.
Latest in Science
Science highlights or most popular articles

Flipping lipids and slime molds
A dull first job nearly pushed JBC associate editor Todd Graham out of science. Then a slime mold project changed his path. Now, he studies membrane biology and reflects on discovery, persistence and mentoring through uncertainty.
How smelling death alters worm behavior
Researchers have found that the roundworm C. elegans can smell death, and it changes how the worms behave, reproduce and age.

A chance encounter with the lab
Payton Stevens never planned to become a pancreatic cancer researcher. A temporary job set him on a path from rural Kentucky to leading research on Wnt signaling and metastasis, where he now pairs discovery with mentorship and science advocacy.

Light-activated small molecule could transform eye infection treatment
Contact lenses raise the risk of infectious keratitis, a leading cause of blindness worldwide. A biotech company is commercializing a light-activated therapy using a ROS-generating molecule to rapidly kill microbes in the cornea to preserve vision.

The molecular orchestra of memory
Calcium, calmodulin and calcium/calmodulin-dependent kinase II form a molecular axis that turns fleeting neural activity into lasting memories. New research shows how memories are stabilized, and possibly even protected or repaired.
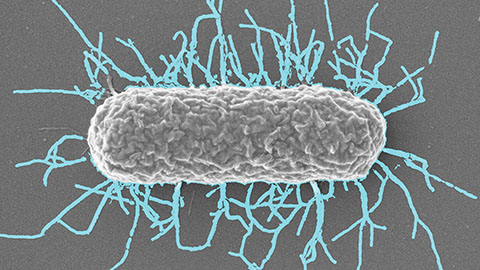
Differences in pili structure modulate bacterial behavior
Researchers demonstrate how small changes in the structure of hair-like protein appendages can affect the behavior of Acinetobacter bacteria.

